fastcpd_lm() and fastcpd.lm() are wrapper
functions of fastcpd() to find change points in linear
regression models. The function is similar to fastcpd() except that
the data is by default a matrix or data frame with the response variable
as the first column and thus a formula is not required here.
Arguments
- data
A matrix or a data frame with the response variable as the first column.
- ...
Other arguments passed to
fastcpd(), for example,segment_count.
Value
A fastcpd object.
Examples
if (requireNamespace("mvtnorm", quietly = TRUE)) {
set.seed(1)
n <- 300
p <- 4
x <- mvtnorm::rmvnorm(n, rep(0, p), diag(p))
theta_0 <- rbind(c(1, 3.2, -1, 0), c(-1, -0.5, 2.5, -2), c(0.8, 0, 1, 2))
y <- c(
x[1:100, ] %*% theta_0[1, ] + rnorm(100, 0, 3),
x[101:200, ] %*% theta_0[2, ] + rnorm(100, 0, 3),
x[201:n, ] %*% theta_0[3, ] + rnorm(100, 0, 3)
)
result_lm <- fastcpd.lm(cbind(y, x))
summary(result_lm)
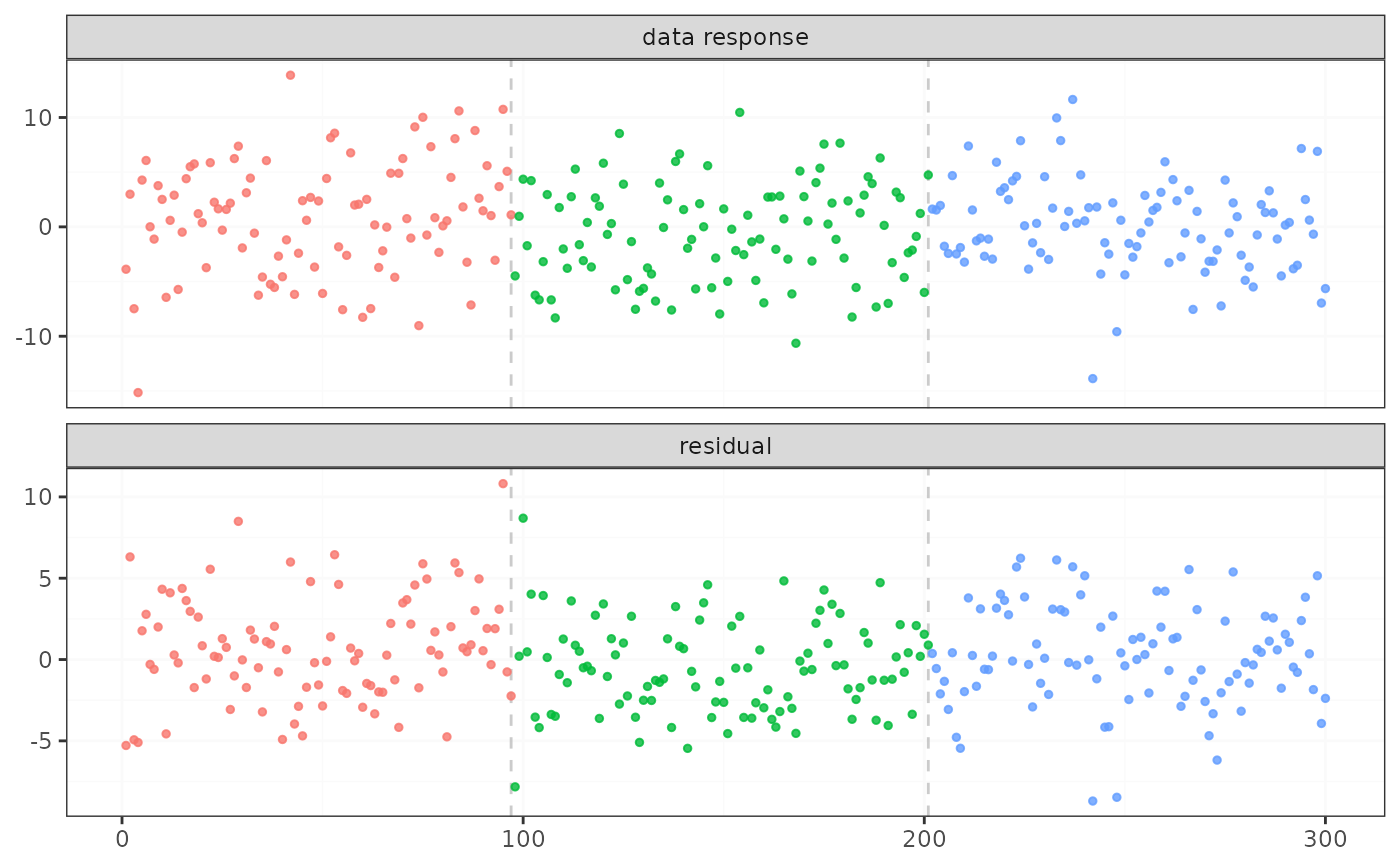
plot(result_lm)
}
#>
#> Call:
#> fastcpd.lm(data = cbind(y, x))
#>
#> Change points:
#> 97 201
#>
#> Cost values:
#> 528.9771 424.3702 471.2645
#>
#> Parameters:
#> segment 1 segment 2 segment 3
#> 1 0.74291290 -0.6153049 0.8733473
#> 2 3.69465275 -0.5034948 0.3222868
#> 3 -1.24746871 2.2522352 1.0188455
#> 4 0.09579985 -1.9875126 2.2761340
 # \donttest{
if (requireNamespace("mvtnorm", quietly = TRUE)) {
set.seed(1)
n <- 600
p <- 4
d <- 2
x <- mvtnorm::rmvnorm(n, rep(0, p), diag(p))
theta_1 <- matrix(runif(8, -3, -1), nrow = p)
theta_2 <- matrix(runif(8, -1, 3), nrow = p)
y <- rbind(
x[1:350, ] %*% theta_1 + mvtnorm::rmvnorm(350, rep(0, d), 3 * diag(d)),
x[351:n, ] %*% theta_2 + mvtnorm::rmvnorm(250, rep(0, d), 3 * diag(d))
)
result_mlm <- fastcpd.lm(cbind.data.frame(y = y, x = x), p.response = 2)
summary(result_mlm)
}
#>
#> Call:
#> fastcpd.lm(data = cbind.data.frame(y = y, x = x), p.response = 2)
#>
#> Change points:
#> 350
#>
#> Cost values:
#> 1431.408 1019.454
#>
#> Parameters:
#> segment 1 segment 2
#> 1 -2.453012 1.68044714
#> 2 -2.295667 -0.46458087
#> 3 -1.327543 1.07765071
#> 4 -2.783358 -0.15196831
#> 5 -1.895117 2.86434615
#> 6 -1.927478 2.61647011
#> 7 -1.168885 1.73783271
#> 8 -1.380168 0.09453771
# }
# \donttest{
if (requireNamespace("mvtnorm", quietly = TRUE)) {
set.seed(1)
n <- 600
p <- 4
d <- 2
x <- mvtnorm::rmvnorm(n, rep(0, p), diag(p))
theta_1 <- matrix(runif(8, -3, -1), nrow = p)
theta_2 <- matrix(runif(8, -1, 3), nrow = p)
y <- rbind(
x[1:350, ] %*% theta_1 + mvtnorm::rmvnorm(350, rep(0, d), 3 * diag(d)),
x[351:n, ] %*% theta_2 + mvtnorm::rmvnorm(250, rep(0, d), 3 * diag(d))
)
result_mlm <- fastcpd.lm(cbind.data.frame(y = y, x = x), p.response = 2)
summary(result_mlm)
}
#>
#> Call:
#> fastcpd.lm(data = cbind.data.frame(y = y, x = x), p.response = 2)
#>
#> Change points:
#> 350
#>
#> Cost values:
#> 1431.408 1019.454
#>
#> Parameters:
#> segment 1 segment 2
#> 1 -2.453012 1.68044714
#> 2 -2.295667 -0.46458087
#> 3 -1.327543 1.07765071
#> 4 -2.783358 -0.15196831
#> 5 -1.895117 2.86434615
#> 6 -1.927478 2.61647011
#> 7 -1.168885 1.73783271
#> 8 -1.380168 0.09453771
# }